studied Biology at the University of Athens, Greece, from 2002 to 2008, where he did his structural biology project in the laboratory of Prof. Stavros Hamodrakas. He performed his PhD work in the field of computational structural biology at the Bijvoet Center for Biomolecular Research, Utrecht, The Netherlands, and obtained his PhD in 2012 under the supervision of Prof. Alexandre Bonvin from the Chemistry Department of Utrecht University. In 2013, he moved to EMBL-Heidelberg to work with Dr. Anne-Claude Gavin and Dr. Martin Beck in the field of structural proteomics with a focus on cryo-electron microscopy. Since August 2018 he is a group leader and Junior Professor at the interdisciplinary research centre ZIK HALOmem at the MLU. His research project focuses on the structural modeling of highly flexible protein-protein interactions using various sources of experimental data (e.g. XL-MS, cryo-EM) to drive the modeling procedure.
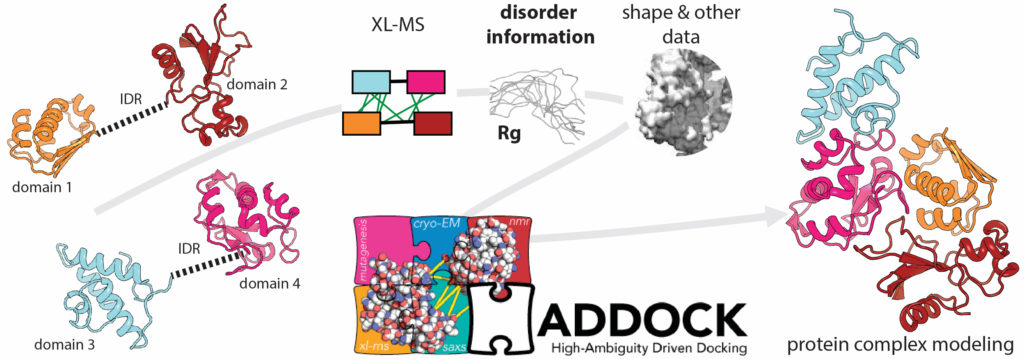
Project within the RTG
Key publications
Kastritis PL, Rodrigues JP, Folkers GE, Boelens R, Bonvin AM. Proteins feel more than they see: fine-tuning of binding affinity by properties of the non-interacting surface. J Mol Biol. 2014, 426: 2632-2652.
Kastritis PL, O’Reilly FJ, Bock T, Li Y, Rogon MZ, Buczak K, Romanov N, Betts MJ, Bui KH, Hagen WJ, Hennrich ML, Mackmull MT, Rappsilber J, Russell RB, Bork P, Beck M, Gavin AC. Capturing protein communities by structural proteomics in a thermophilic eukaryote. Mol Syst Biol. 2017, 13: 936.
van Zundert GCP, Rodrigues JPGLM, Trellet M, Schmitz C, Kastritis PL, Karaca E, Melquiond ASJ, van Dijk M, de Vries SJ, Bonvin AM. The HADDOCK2.2 web server: user-friendly integrative modeling of biomolecular complexes. J Mol Biol. 2016, 428: 720-725.
Website: https://www.halomem.de/en/research-group-kastritis/#forschung